

From genetic discovery to cardiovascular innovation
Lab Mission The Weldy Lab seeks to transform cardiovascular medicine by integrating human genetics with mechanistic biology to uncover causal pathways of disease and pioneer new therapeutic strategies.
Defining genetic mechanisms of cardiovascular disease

The Weldy Lab studies how human genetic variation drives vascular dysfunction, inflammation, and cardiovascular disease. We integrate human genetics, epigenetics, single-cell genomics, RNA biology, and experimental models to define causal mechanisms and identify new therapeutic opportunities.
Led by Chad S. Weldy, MD, PhD, the lab focuses on questions at the intersection of cardiovascular genetics, vascular biology, and RNA-mediated regulation, with particular interest in smooth muscle cell phenotypic modulation, innate immune signaling, and mechanisms of disease susceptibility governed by epigenetic memory.
Our work is grounded in the idea that genetic discovery can do more than identify risk. It can reveal the biology that matters most and point toward more precise strategies for prevention and treatment.
Featured publications
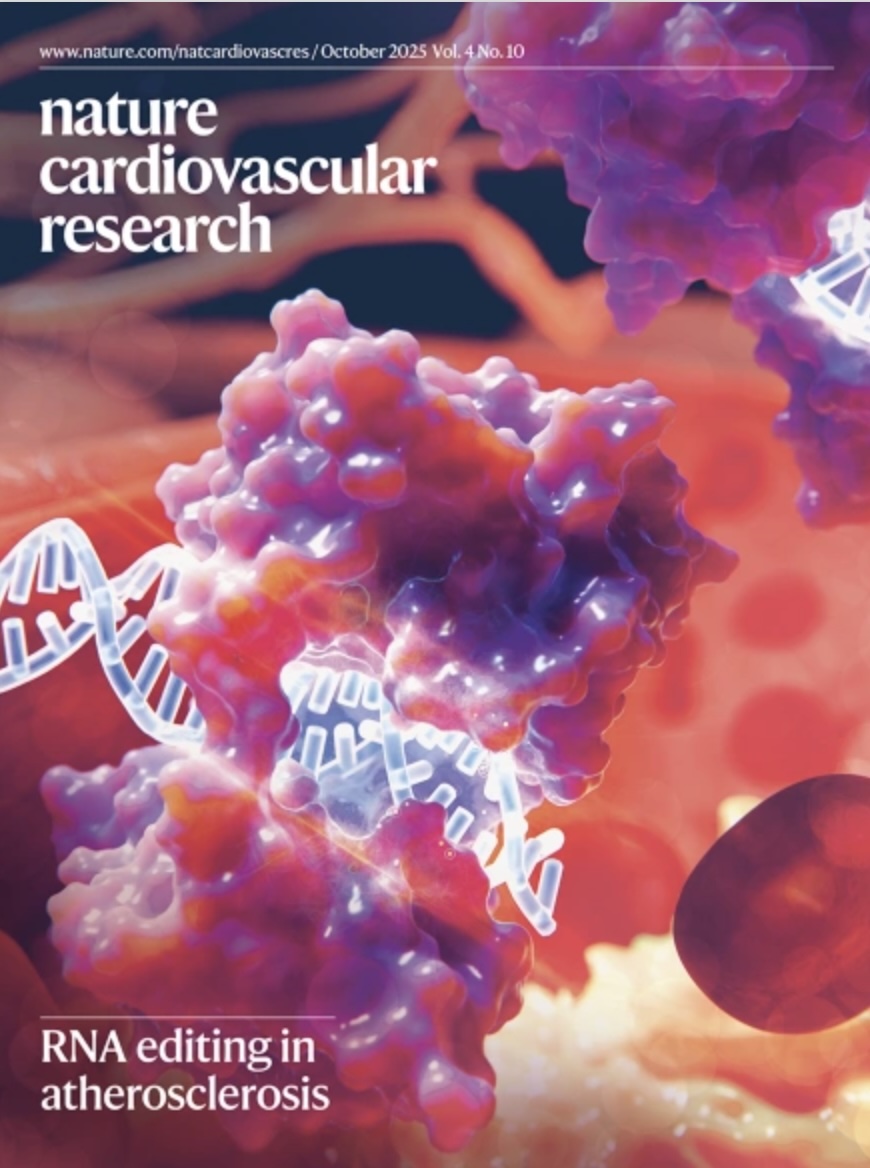
Smooth muscle cell ADAR1 controls activation of RNA sensor MDA5 in atherosclerosis
Weldy CS, et al. Nature Cardiovascular Research (2025)
DOI: 10.1038/s44161-025-00710-5
A smooth muscle–specific RNA editing program regulates dsRNA sensing and inflammatory signaling in atherosclerosis, linking RNA biology to vascular disease mechanisms.
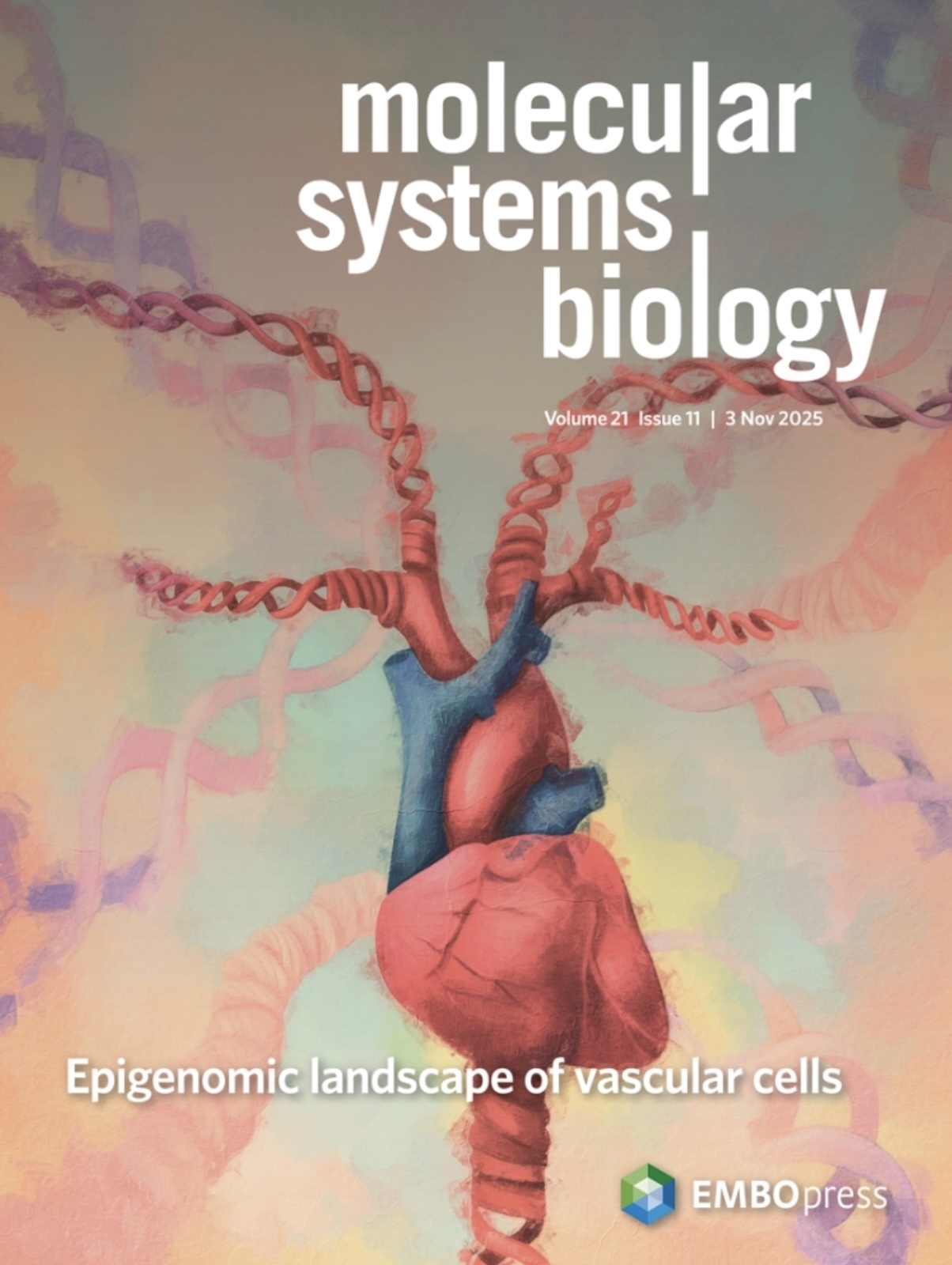
Epigenomic landscape of single vascular cells reflects developmental origin and disease risk loci
Weldy CS, et al. Molecular Systems Biology (2025)
DOI: 10.1038/s44320-025-00140-2
Single-cell epigenomic profiling reveals how developmental origin shapes vascular cell states and maps aortic dimension and aneurysm risk loci to specific cell type and vascular site specific programs across smooth muscle, endothelial, and fibroblast cell types.
Areas of focus
- Human genetics & causality
- Vascular biology & site-specific mechanisms
- Epigenetics and epigenetic memory
- RNA biology (A-to-I editing) and innate immune sensing
- Single-cell and multi-omics approaches
RNA editing and double-stranded RNA sensing
RNA editing by ADAR enzymes is a fundamental mechanism that allows cells to distinguish self from non-self RNA.
Through A-to-I editing of double-stranded RNA, ADAR1 reshapes endogenous RNA structures and prevents inappropriate activation of the innate immune sensor MDA5 (gene symbol IFIH1).
Genetic and molecular studies now show that reduced RNA editing increases inflammatory signaling and disease risk, including coronary artery disease. Our work demonstrates that impaired ADAR1-mediated editing in vascular smooth muscle cells leads to pathologic dsRNA sensing, identifying endogenous RNA recognition as a causal mechanism of vascular disease.
By integrating human genetics, RNA biology, and vascular models, we study how variation in RNA editing and activation of dsRNA sensing programs connects genetic risk to inflammation and cardiovascular pathology.
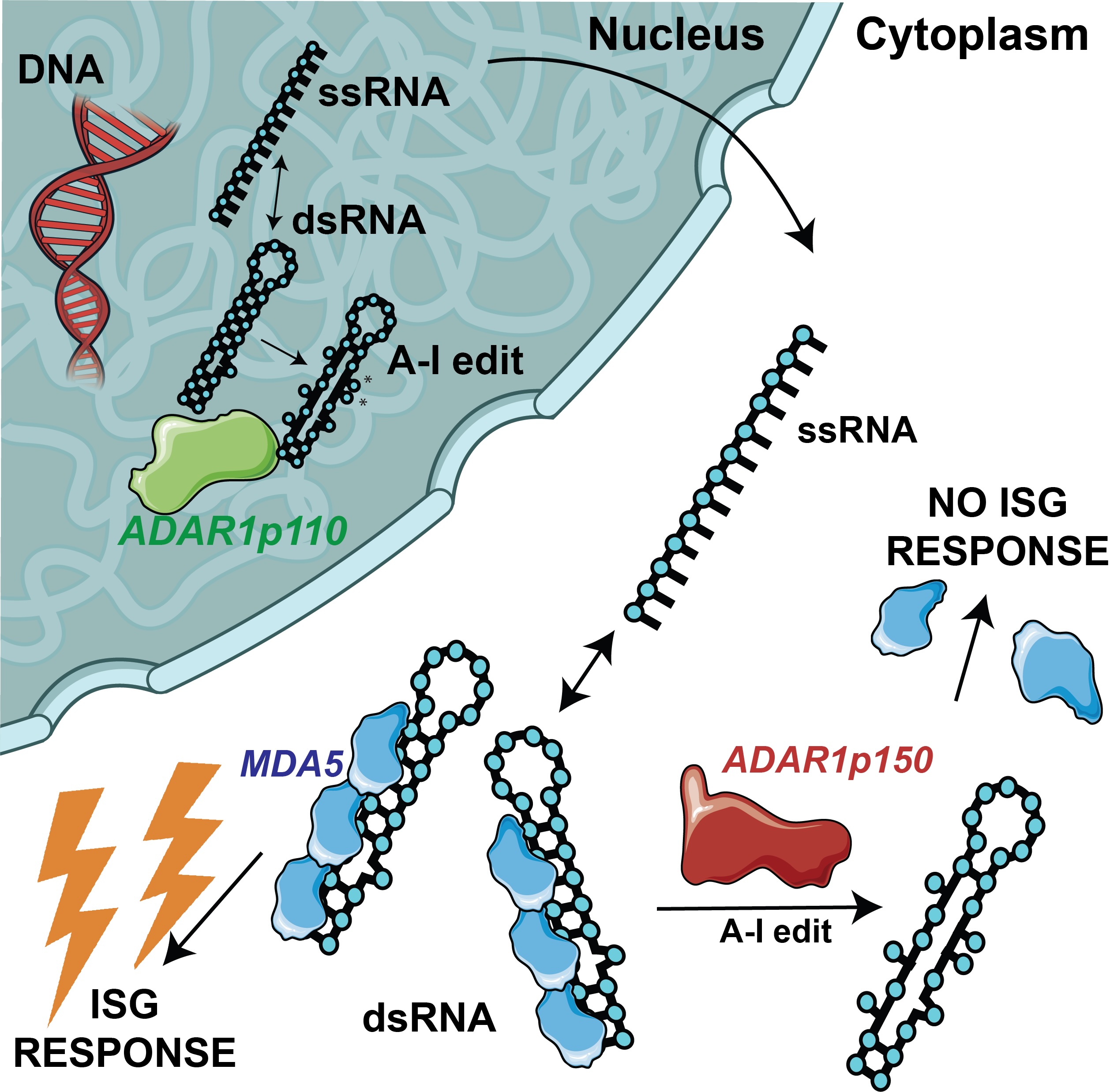 ADAR1-mediated RNA editing suppresses endogenous dsRNA sensing by MDA5 in vascular cells.
ADAR1-mediated RNA editing suppresses endogenous dsRNA sensing by MDA5 in vascular cells.
Recently reviewed by Weldy et al. ATVB, 2026 Read the paper →
News
Stanford Medicine highlights our RNA editing work
A Stanford Medicine feature on ADAR1 RNA editing, dsRNA sensing, and implications for cardiovascular disease mechanisms.
Read the story →
Stanford CVI story on developmental “memory” in arteries
How developmental origin leaves a lasting imprint shaping regional vascular disease risk.
Read the story →
Affiliations




Funding